
- The basic input data is inserted in the CTRL file.
The CTRL file shown below contains the minimum amount of
information to be typed, in order to run the TB-LMTO programs. The rest
may be obtained from the default values in the program, which are
written into the CTRL file by the program.
HEADER NiSe2 Pyrit (Acta Chem. Scand. v. 23 p. 2325 1969) VERS LMASA-47 STRUC ALAT=11.2683 PLAT=1 0 0 0 1 0 0 0 1 CLASS ATOM=NI Z=28 ATOM=Se Z=34 SITE ATOM=NI POS= .0 .0 .0 ATOM=NI POS= .5 .0 .5 ATOM=NI POS= .0 .5 .5 ATOM=NI POS= .5 .5 .0 ATOM=Se POS= .383 .383 .383 ATOM=Se POS= .617 .617 .617 ATOM=Se POS= .117 .617 .883 ATOM=Se POS= .883 .383 .117 ATOM=Se POS= .617 .883 .117 ATOM=Se POS= .383 .117 .883 ATOM=Se POS= .883 .117 .617 ATOM=Se POS= .117 .883 .383The input consists of tokens, like ATOM= or Z=. These are grouped together in categories, like CLASS. The name of a category always starts in the first column of a line. This will be explained in detail in Section V. ALAT= is a lattice constant in a.u. (i.e. Bohr radii), PLAT= are the lattice translation vectors and POS= are the atomic positions. The latter two are in Cartesian coordinates and in units of ALAT.
All Ni atoms and all Se atoms are equivalent i.e. there are two classes. From the information in the CTRL file, the program will find the symmetry i.e. all space group operations and the generators of the space group. If the symmetry of the bravais lattice is incompatible with the symmetry of the basis, then the symmetry of the bravais lattice is used. Alternatively to the CTRL file above, the symmetry generators and only one of each of the inequivalent atoms could have been given. The program will complete the basis:
HEADER NiSe2 Pyrit (Acta Chem. Scand. v. 23 p. 2325 1969) VERS LMASA-47 STRUC ALAT=11.2683 PLAT=1 0 0 0 1 0 0 0 1 CLASS ATOM=NI Z=28 ATOM=Se Z=34 SITE ATOM=NI POS= .0 .0 .0 ATOM=Se POS= .383 .383 .383 SYMGRP NGEN=3 GENGRP=I R3D MX:(.5,.5,0)The category SYMGRP contains three tokens: NGEN= the number of generators supplied,
GENGRP= the names of the generators as defined in the Subsection `Category SYMGRP'.A third possibility is to use the interactive program lminit.run in the main directory. An example is shown below, where the user's answer to the query is indicated by the string after `->' and followed by `(Answer)'.
| QUERY : SPCGRP: | SYMBOL OR NUMBER OF SPACE GROUP | | Enter: 'nnn', nnn is the value for SPCGRP, | 'a' to abort program execution. | ->205 (Answer) Space group: Pa-3 No.:205 Crystal system: cubic Generators: I R3D MX::(1/2,1/2,0) | QUERY : atomic-units ? (else in Angstrom) | | Enter: 'nnn', nnn is the value for ATUNITS, | default value=T | 'a' to abort program execution. | '/' to accept default value | ->/ Now enter lattice parameters | QUERY : A:lattice constant (atomic units) | | Enter: 'nnn', nnn is the value for A, | 'a' to abort program execution. | ->11.2683 (Answer) CTRLUC: PLAT ALAT=11.2683 1.00000 .00000 .00000 .00000 1.00000 .00000 .00000 .00000 1.00000 | QUERY: label or nuclear charge of class 1 | | Enter: 'nnn', nnn is the value for label, | 'a' to abort program execution. | 'q' to quit | ->28 (Answer) Class: Ni ; nuclear charge 28 (atom Ni) | QUERY: X = position of Ni (in units of the | translation vectors a, b, c) | | Enter: 'nnn', nnn is the value for X, | default value=0. 0. 0. | 'a' to abort program execution. | '/' to accept default value | ->/ (Answer) | QUERY: label or nuclear charge of class 2 | | Enter: 'nnn', nnn is the value for label, | 'a' to abort program execution. | 'q' to quit | ->34 (Answer) Class: Se ; nuclear charge 34 (atom Se) | QUERY : X = position of Se | | Enter: 'nnn', nnn is the value for X, | default value=0. 0. 0. | 'a' to abort program execution. | '/' to accept default value | ->.383 .383 .383 | QUERY: label or nuclear charge of class 3 | | Enter: 'nnn', nnn is the value for label, | 'a' to abort program execution. | 'q' to quit | ->q (Answer) CHKSYM: Check the symmetry of the crystal: GENGRP: A subgroup of the space group will be created from 3 generators The generators created 24 sym. ops. ADDBAS: The basis has been extended from 2 to 12 atoms. The new positions are: ATOM=Ni POS= .500 .500 .000 ATOM=Ni POS= .000 .500 .500 ATOM=Ni POS= .500 .000 .500 ATOM=Se POS= -.383 -.383 -.383 ATOM=Se POS= .117 .883 .383 ATOM=Se POS= .883 .117 -.383 ATOM=Se POS= .383 .117 .883 ATOM=Se POS= .883 .383 .117 ATOM=Se POS= -.383 .883 .117 ATOM=Se POS= .117 -.383 .883 SYMLAT: lattice invariant under 48 sym. ops. SYMCRY: crystal invariant under 24 sym. ops. SHOSYM: Bravais system : cubic Bravais lattice: cubic primitive Centring type : P Crystal family : cubic Crystal system : cubic Point group : m-3The resulting CTRL file looks like this:
HEADER Ni4Se8, cubic primitive SYMGRP NGEN=3 GENGRP=I R3D MX:(.5,.5,0) SPCGRP=Pa-3 STRUC ALAT=11.2683 PLAT=1 0 0 0 1 0 0 0 1 CLASS ATOM=Ni Z=28 ATOM=Se Z=34 SITE ATOM=Ni POS=0.000 0.000 0.000 ATOM=Ni POS=0.500 0.500 0.000 ATOM=Ni POS=0.500 0.000 0.500 ATOM=Ni POS=0.000 0.500 0.500 ATOM=Se POS=0.383 0.117 -.117 ATOM=Se POS=-.383 -.117 0.117 ATOM=Se POS=-.117 0.383 0.117 ATOM=Se POS=0.117 -.383 -.117 ATOM=Se POS=0.117 -.117 0.383 ATOM=Se POS=-.117 0.117 -.383 ATOM=Se POS=0.383 0.383 0.383 ATOM=Se POS=-.383 -.383 -.383 - The next step is to find the size of the atomic spheres.
First muffin-tin (MT) radii are found by lmhart.run and inserted
into the CTRL file. The result of this run is shown below.
HEADER Ni4Se8, cubic primitive VERS LMASA-47 IO VERBOS=30 OUTPUT=LM ERR=ERR SYMGRP NGEN=3 GENGRP=I R3D MX:(.5,.5,0) SPCGRP=Pa-3 STRUC ALAT=11.2683 PLAT=1 0 0 0 1 0 0 0 1 DIM NBAS=12 NCLASS=2 NL=3 LDIM=68 IDIM=40 NSYMOP=24 CLASS ATOM=Ni Z=28 R=2.41776510 ATOM=Se Z=34 R=2.28352037 SITE ATOM=Ni POS=0.000 0.000 0.000 ATOM=Ni POS=0.500 0.500 0.000 ATOM=Ni POS=0.500 0.000 0.500 ATOM=Ni POS=0.000 0.500 0.500 ATOM=Se POS=0.383 0.117 -.117 ATOM=Se POS=-.383 -.117 0.117 ATOM=Se POS=-.117 0.383 0.117 ATOM=Se POS=0.117 -.383 -.117 ATOM=Se POS=0.117 -.117 0.383 ATOM=Se POS=-.117 0.117 -.383 ATOM=Se POS=0.383 0.383 0.383 ATOM=Se POS=-.383 -.383 -.383The two tokens (R=) with the MT-radii in a.u. have been inserted in category CLASS. Furthermore, a new class, IO, has been inserted with the tokens: VERBOS= which specifies the amount of output the programs print and the number of default tokens which will be inserted in the CTRL file by the programs which updates the CTRL file, OUTPUT= which gives a file name to which standard out will be directed, ERR= which gives a file name to which error messages will be directed.
- The overlap between the WS-spheres is checked by the
lmovl.run program.
The result written to standard out
looks like (the output has been edited to fit into this
documentation):
Info: impossible to reach VOL, increase OMMAX. CELL VOLUME= 1430.78771, SUM OF SPHERE VOLUMES= 992.45473 IB JB CL1 CL2 DIST SUMRS 1/d 1/s1 1/s2 ========================================== 1 5 Ni Se .42 .48 16 27 28 ------------------------------------------ 5 1 Se Ni .42 .48 16 28 27 5 6 Se Se .41 .47 16 28 28 ------------------------------------------
The first line tells that space filling could not be reached with the default maximum overlaps (two atomic spheres are allowed to overlap no more than 16% in the sence of 1/d below, max. overlap between an atomic and an interstitial sphere is 18%, and max. overlap between two interstitial spheres is 20%).
The next two lines show the cell volume and the volume inside the expanded spheres.
In the table, each line displays two neighboring atoms with WS-radii s1 and s2, and the distance between them d. Under `1/d', `1/s1', and `1/s2' are the results of
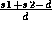 ,
, 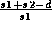 , and
, and
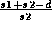 , respectively, in percent.
If the two sphere radii are very much different, one should check
`1/s1', where s1 is the smallest radius. (A big sphere may swallow a
small sphere.)
, respectively, in percent.
If the two sphere radii are very much different, one should check
`1/s1', where s1 is the smallest radius. (A big sphere may swallow a
small sphere.)
Since in the present case space filling could not be reached, interstitial spheres have to be inserted.
- Interstitial spheres are found by the lmes.run program.
The resulting CTRL file is shown below.
HEADER Ni4Se8, cubic primitive VERS LMASA-47 IO VERBOS=30 OUTPUT=OVL ERR=ERR SYMGRP NGEN=3 GENGRP=I R3D MX:(.5,.5,0) SPCGRP=Pa-3 STRUC ALAT=11.2683 PLAT=1 0 0 0 1 0 0 0 1 DIM NBAS=36 NCLASS=3 NL=3 LDIM=68 IDIM=40 NSYMOP=24 CLASS ATOM=Ni Z=28 R=2.41776510 ATOM=Se Z=34 R=2.28352037 ATOM=E Z= 0 R=1.53150056 SITE ATOM=Ni POS=0.00000 0.00000 0.00000 ATOM=Ni POS=0.50000 0.50000 0.00000 ATOM=Ni POS=0.50000 0.00000 0.50000 ATOM=Ni POS=0.00000 0.50000 0.50000 ATOM=Se POS=0.38300 0.11700 -.11700 ATOM=Se POS=-.38300 -.11700 0.11700 ATOM=Se POS=-.11700 0.38300 0.11700 ATOM=Se POS=0.11700 -.38300 -.11700 ATOM=Se POS=0.11700 -.11700 0.38300 ATOM=Se POS=-.11700 0.11700 -.38300 ATOM=Se POS=0.38300 0.38300 0.38300 ATOM=Se POS=-.38300 -.38300 -.38300 ATOM=E POS=0.28510 -.18060 0.09608 ATOM=E POS=-.28510 0.18060 -.09608 ATOM=E POS=0.09608 0.28510 -.18060 ATOM=E POS=-.09608 -.28510 0.18060 ATOM=E POS=0.18060 -.09608 -.28510 ATOM=E POS=-.18060 0.09608 0.28510 ATOM=E POS=0.31939 0.09608 0.21489 ATOM=E POS=0.21489 0.31939 0.09608 ATOM=E POS=-.31939 -.09608 -.21489 ATOM=E POS=-.21489 -.31939 -.09608 ATOM=E POS=0.09608 0.21489 0.31939 ATOM=E POS=-.09608 -.21489 -.31939 ATOM=E POS=0.40391 -.21489 -.18060 ATOM=E POS=-.40391 0.21489 0.18060 ATOM=E POS=-.18060 0.40391 -.21489 ATOM=E POS=0.18060 -.40391 0.21489 ATOM=E POS=0.21489 0.18060 -.40391 ATOM=E POS=-.21489 -.18060 0.40391 ATOM=E POS=0.40391 -.28510 0.31939 ATOM=E POS=0.31939 0.40391 -.28510 ATOM=E POS=-.40391 0.28510 -.31939 ATOM=E POS=-.31939 -.40391 0.28510 ATOM=E POS=-.28510 0.31939 0.40391 ATOM=E POS=0.28510 -.31939 -.40391The program found 24 interstitial spheres, all equivalent, with a sphere radius between 0.9 and 4.0 a.u. These limits are default and can be changed. Notice that a new category DIM has been inserted, which contain information about some dimensions.
- Check the overlap again with lmovl.run. Standard out now
looks like:
IB JB CL1 CL2 DIST SUMRS 1/d 1/s1 1/s2 ========================================= 1 5 NI Se .42 .47 13 22 23 1 20 NI E1 .35 .40 13 18 29 ----------------------------------------- 5 1 Se NI .42 .47 13 23 22 5 6 Se Se .41 .46 13 23 23 5 14 Se E1 .34 .38 13 19 28 5 15 Se E1 .38 .38 1 1 2 5 17 Se E1 .34 .38 13 19 28 5 22 Se E1 .34 .38 13 19 28 5 23 Se E1 .34 .38 13 19 28 5 29 Se E1 .38 .38 1 1 2 5 31 Se E1 .34 .38 13 19 28 5 34 Se E1 .34 .38 13 19 28 5 35 Se E1 .38 .38 1 1 2 5 36 Se E1 .34 .38 13 19 28 ----------------------------------------- 13 2 E1 NI .35 .40 13 29 18 13 6 E1 Se .34 .38 13 28 19 13 7 E1 Se .38 .38 1 2 1 13 8 E1 Se .34 .38 13 28 19 13 12 E1 Se .34 .38 13 28 19 13 15 E1 E1 .27 .31 12 21 21 13 25 E1 E1 .30 .31 1 2 2 -----------------------------------------
The program found space filling with a maximum overlap of 13% which is fine and one can proceed.
- Complete the CTRL file with the lmctl.run program, but first
change the token VERBOS= to 50 in category IO to ensure that all
default values are inserted.
HEADER Ni4Se8, cubic primitive VERS LMASA-47 IO VERBOS=50 HELP=F WKP=F IACTIV=F ERRTOL=2 OUTPUT=CTL ERR=ERR SYMGRP NGEN=3 GENGRP=I R3D MX:(.5,.5,0) SPCGRP=Pa-3 USESYM=F STRUC ALAT=11.2683 PLAT=1 0 0 0 1 0 0 0 1 FIXLAT=T DIM NBAS=36 NCLASS=3 NL=3 LDIM=92 IDIM=232 NSYMOP=24 NKP=11 OPTIONS NSPIN=1 REL=T CCOR=T NONLOC=F NRXC=1 NRMIX=2 CORDRD=F NITATOM=30 CHARGE=F FATBAND=F AFM=F FS=F CARTESIAN=T WRIBAS=F Q=---- CLASS ATOM=Ni Z=28 R=2.72719158 LMX=2 CONF=4 4 3 4 IDXDN=1 1 1 IDMOD=0 0 0 ATOM=Se Z=34 R=2.57576616 LMX=2 CONF=4 4 4 4 IDXDN=1 1 2 IDMOD=0 0 0 ATOM=E Z= 0 R=1.72750257 LMX=2 CONF=1 2 3 4 IDXDN=1 2 2 IDMOD=0 0 0 SITE ATOM=Ni POS=0.00000 0.00000 0.0000 ATOM=Ni POS=0.50000 0.50000 0.0000 ATOM=Ni POS=0.50000 0.00000 0.5000 ATOM=Ni POS=0.00000 0.50000 0.5000 ATOM=Se POS=0.38300 0.11700 -.1170 ATOM=Se POS=-.38300 -.11700 0.1170 ATOM=Se POS=-.11700 0.38300 0.1170 ATOM=Se POS=0.11700 -.38300 -.1170 ATOM=Se POS=0.11700 -.11700 0.3830 ATOM=Se POS=-.11700 0.11700 -.3830 ATOM=Se POS=0.38300 0.38300 0.3830 ATOM=Se POS=-.38300 -.38300 -.3830 ATOM=E POS=0.28510 -.18060 0.0960 ATOM=E POS=-.28510 0.18060 -.0960 ATOM=E POS=0.09608 0.28510 -.1806 ATOM=E POS=-.09608 -.28510 0.1806 ATOM=E POS=0.18060 -.09608 -.2851 ATOM=E POS=-.18060 0.09608 0.2851 ATOM=E POS=0.31939 0.09608 0.2148 ATOM=E POS=0.21489 0.31939 0.0960 ATOM=E POS=-.31939 -.09608 -.2148 ATOM=E POS=-.21489 -.31939 -.0960 ATOM=E POS=0.09608 0.21489 0.3193 ATOM=E POS=-.09608 -.21489 -.3193 ATOM=E POS=0.40391 -.21489 -.1806 ATOM=E POS=-.40391 0.21489 0.1806 ATOM=E POS=-.18060 0.40391 -.2148 ATOM=E POS=0.18060 -.40391 0.2148 ATOM=E POS=0.21489 0.18060 -.4039 ATOM=E POS=-.21489 -.18060 0.4039 ATOM=E POS=0.40391 -.28510 0.3193 ATOM=E POS=0.31939 0.40391 -.2851 ATOM=E POS=-.40391 0.28510 -.3193 ATOM=E POS=-.31939 -.40391 0.2851 ATOM=E POS=-.28510 0.31939 0.4039 ATOM=E POS=0.28510 -.31939 -.4039 SCALE SCLWSR=T OMMAX1=.16 .18 .20 OMMAX2=.40 .45 .50 STR KAPPA2=0 RMAXS=3.2 NDIMIN=350 NOCALC=F IALPHA=0 DOWATS=F DELTR=.1 LMAXW=8 ATOM=Ni SIGMA=.7 .7 .7 ATOM=Se SIGMA=.7 .7 .7 ATOM=E SIGMA=.7 .7 .7 START NIT=30 BROY=T WC=-1 NMIX=1 BETA=.5 FREE=F CNVG=.00001 CNVGET=.00001 BEGMOM=T CNTROL=T EFERMI=-.25 VMTZ=-.75 ATOM=Ni P=4.67 4.41 3.86 Q=0.6 0.0 0.0 0.8 0.0 0.0 8.6 0.0 0.0 enu =-.543 -.398 -.304 c =-.419 0.544 -.281 sqrdel=0.464 0.580 0.188 p =0.054 0.027 2.398 gamma =0.546 0.240 -.009 ATOM=Se P=4.95 4.84 4.17 Q=2.0 0.0 0.0 3.7 0.0 0.0 0.3 0.0 0.0 enu =-1.005 -0.388 -0.513 c =-1.304 -0.360 1.311 sqrdel= 0.344 0.387 0.596 p = 0.372 0.170 0.024 gamma = 0.442 0.145 0.150 ATOM=E P=1.5 2.5 3.5 Q=0 0 0 0 0 0 0 0 0 enu =-1.758 -0.642 1.316 c = 0.990 3.503 6.985 sqrdel= 0.590 0.485 0.400 p = 0.006 0.003 0.001 gamma = 0.358 0.065 0.022 CHARGE LMTODAT=T ELF=F ADDCOR=F SPINDENS=F CHARWIN=F EMIN=-2 EMAX=2 PLOT ORIGIN=0 0 0 R1=1 0 0 NDELR1=0 R2=0 1 0 NDELR2=0 R3=0 0 1 NDELR3=0 FORMAT=1 BZ NKABC=4 4 4 TETRA=T METAL=T TOL=.000001 N=0 W=.005 RANGE=5 NPTS=1001 EWALD NKDMX=250 AS=2 TOL=.000001 RHOFIT FIT=F KAPPA2=0 RMAXS=3.5 ATOM=Ni LMXRHO=2 SIGMA=.7 .7 .7 ATOM=Se LMXRHO=2 SIGMA=.7 .7 .7 ATOM=E LMXRHO=2 SIGMA=.7 .7 .7 SCELL PLAT=1 0 0 0 1 0 0 0 1 EQUIV=T HARTREE BEGATOM=T LT1=2 LT2=2 LT3=2 DOS NOPTS=801 EMIN=-2 EMAX=2 SYML NQ=30 Q1=0.0 0.0 0.0 LAB1=g Q2=0.5 0.5 0.0 LAB2=M NQ=20 Q1=0.5 0.5 0.0 LAB1=M Q2=0.5 0.0 0.0 LAB2=X NQ=20 Q1=0.5 0.0 0.0 LAB1=X Q2=0.0 0.0 0.0 LAB2=g NQ=35 Q1=0.0 0.0 0.0 LAB1=g Q2=0.5 0.5 0.5 LAB2=R FINDES RMINES=.9 RMAXES=4 NRXYZ=72 72 72Notice that the Wigner-Sitz radii have been inserted because SCLWSR=T.
This is a complete CTRL file. The meaning of only a few of these tokens is necessary in order to run the program.
7. Before calculating the TB real space structure constants
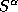 and
and 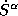 for the combined correction term by the
lmstr.run program, the verbosity is lowered by changing
VERBOS= to 40 in category IO.
for the combined correction term by the
lmstr.run program, the verbosity is lowered by changing
VERBOS= to 40 in category IO.
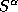 only depends on the structure
and the number of partial waves included, while
only depends on the structure
and the number of partial waves included, while 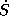 also depends on
the WS-radii. If these are changed during the calculations, the
structure constants have to be recalculated.
also depends on
the WS-radii. If these are changed during the calculations, the
structure constants have to be recalculated.

0 Comments